CONTACT
Department of Chemistry
Todd Wehr Chemistry, 101
1414 W Clybourn St
PO Box 1881
Milwaukee, WI 53233
(414) 288-3515
Problem with this webpage?
Report an accessibility problem
Any other problem, contact muchem@marquette.edu.
 Dr. Nicholas ReiterMarquette University
Dr. Nicholas ReiterMarquette UniversityWehr Chemistry Building, 570
MilwaukeeWI53201United States of America(414) 288-5701nicholas.reiter@marquette.eduThe Reiter LabPh.D., University of Wisconsin-Madison
B.A., Carleton College
Ribonucleic acid (RNA) molecules play a central role in life, coordinating intertwined layers of regulation that extend far beyond its role as a messenger. RNA is ideally suited to orchestrate this biological diversity because it is the only macromolecule that can both carry genetic information and perform chemical reactions by acting as an enzyme. We are at an exciting frontier and are beginning to comprehend how RNA molecules interact with macromolecular protein complexes and how RNA tertiary structures function in biology.
Our mission is to prepare the next generation of scientists and advance our understanding of RNA-protein interactions and structural biology by:
1) Understanding the diverse roles of RNA tertiary structure in biology
2) Defining the conformational motions that enable RNAs to perform shape-based molecular recognition.
3) Elucidating novel mechanisms of RNA-chromatin-modifying enzyme interactions.
To investigate the function and conformational dynamics of RNA-protein (RNP) interactions, we combine quantitative biophysical tools (including X-ray crystallography, nuclear magnetic resonance (NMR) spectroscopy, and electron paramagnetic resonance (EPR) spectroscopy) with cell-based biochemical methods.
Current projects in the Reiter group center on an R01-NIH sponsored project to examine the RNA-mediated mechanisms of the lysine specific histone demethylase-1 (LSD1 or KDM1A) enzyme, a flavin-dependent amine oxidase that plays a key role in embryogenesis and epigenetic “reprogramming” of mammalian cells. Importantly, LSD1 has known roles in cell differentiation in cancer, serving as a predictor of metastatic tumors (in prostate, neuroendocrine, and estrogen receptor (ER)(-) breast cancer).
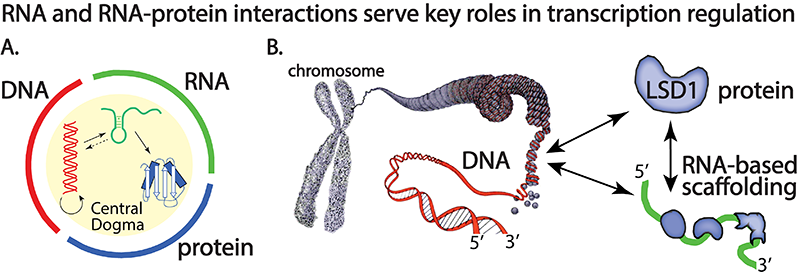
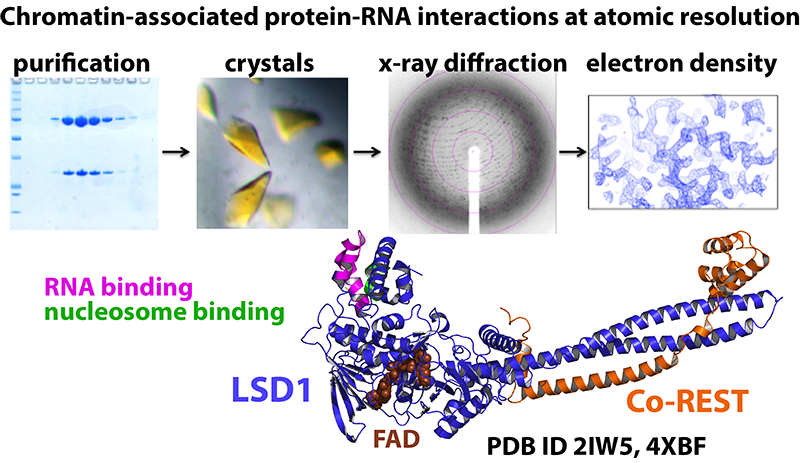
Due to LSD1’s diverse gene regulatory network, novel ways to probe LSD1 in a pathway-specific manner has high therapeutic value. We use the RNA-LSD1 network as model system to examine chromatin-associated protein - RNA interactions at atomic resolution. To this end, we seek to identify and characterize novel RNA-LSD1 interactions throughout the human genome and use this information to develop useful therapeutics that selectively target LSD1 mediated biochemical processes.
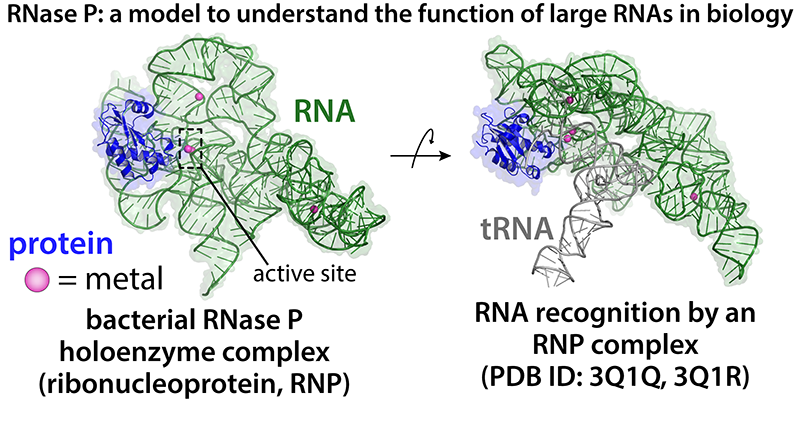
A second project in the lab is to use the RNA-protein enzyme, Ribonuclease P (RNase P), as a model to understand how large RNAs and RNA binding proteins contribute to catalysis. RNase P is required during transfer RNA (tRNA) biogenesis from bacteria to humans, and represents a nearly universal processing event that is linked to the origins of the genetic code. The discovery that the RNA component of RNase P is directly involved in catalysis demonstrated that RNA, just like proteins, is a highly versatile biomolecule and can assemble into a complex three-dimensional architecture. RNase P serves as an ideal system to explore how proteins assemble and interact with large RNA biomolecules. For this project, we integrate both biochemistry and biophysical methods, focusing on the conformational dynamics of RNase P at atomic resolution.
Dr. Nicholas (Nick) Reiter received his B.A. in chemistry from Carleton College, and subsequently performed biochemistry research at the Mayo Clinic (MN). As a graduate student (2001-2006), he was introduced to the RNA world and applied NMR spectroscopy and biophysical methods to understand components involved in the assembly and activation of the spliceosome under the direction of Professor Sam Butcher. He was an NIH post-doctoral fellow (2007-2011) in Professor Alfonso Mondragon’s laboratory at Northwestern University and developed a strong interest in elucidating the tertiary structure of large catalytic RNAs using x-ray crystallography. In 2012, Dr. Reiter joined the faculty at Vanderbilt where he began his NIH funded research program to understand the mechanisms of RNA-chromatin-remodeling enzyme interactions. He joined Marquette in the fall of 2017 and his primary interests are in the area of RNA-protein structure and function.

Reiter lab at Kopp's frozen custard (Summer 2018)