Emeriti & Former Faculty
CONTACT
Department of Biological Sciences
Wehr Life Sciences, 109
1428 W. Clybourn St.
Milwaukee, WI 53233
(414) 288-7355
 Dr. Chris W. MarshallMarquette University
Dr. Chris W. MarshallMarquette UniversityWehr Life Sciences, 511
MilwaukeeWI53201United States of America(414) 288-1698christopher.marshall@marquette.eduMarshall Lab Websitee-PublicationsB.S. 2006, Virginia Tech, Blacksburg, VA
Ph.D. 2013, Medical University of South Carolina, Charleston, SC
Postdoctoral Fellow, 2013-2016, Argonne National Lab/University of Chicago, Chicago, IL
Research Assistant Professor, 2016-2019, University of Pittsburgh, PA
BIOL 4802: Experimental Microbiology
My research program addresses two grand challenges in science: antimicrobial resistance and quantifying global biogeochemical cycling. In the first, I am interested in how diversity and metabolic changes in biofilms contribute to the alarming and expanding problem of antibiotic resistance. In the second, I study anaerobic metabolisms associated with biogeochemical cycles as a biotechnological and ecological source of innovation. Traditionally these disciplines don’t intersect. I would argue, however, that microbial ecology and evolution mediated by metabolic feedback lie at the core of each. Metabolic byproducts can open up new niches that influence eco-evolutionary dynamics and vice versa. Insights into diversification, resilience, resource competition, and cooperation in one system (environment) can inform another (host). Furthermore, the practical component uniting my research program is the study of ecology and evolution in anaerobic biofilms, particularly those that experience limiting resources (e.g. mucosal surfaces or natural environments). In addition, I am keenly interested in developing novel engineered devices and bioinformatic tools to address key scientific questions.
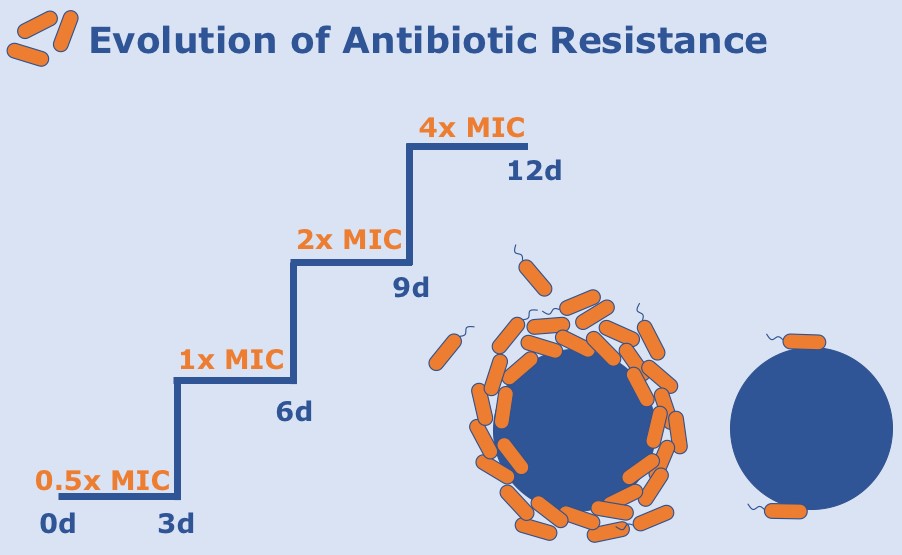 Antibiotic tolerance is the transcriptional/translational change in metabolism that allows for microorganisms to persist in the presence of antibiotics. Antibiotic resistance, on the other hand, is the heritable changes in genomes that render the antibiotics ineffective. I am interested in using high-throughput, automated serial passaging innovations to study antimicrobial tolerance and resistance in biofilm populations. Specifically, I am interested in multi-strain and species interactions that alter antibiotic resistance outcomes. In addition to microbial community dynamics in the presence of antibiotics, I explore the interplay between tolerance and resistance that arises in anaerobic biofilms.
Antibiotic tolerance is the transcriptional/translational change in metabolism that allows for microorganisms to persist in the presence of antibiotics. Antibiotic resistance, on the other hand, is the heritable changes in genomes that render the antibiotics ineffective. I am interested in using high-throughput, automated serial passaging innovations to study antimicrobial tolerance and resistance in biofilm populations. Specifically, I am interested in multi-strain and species interactions that alter antibiotic resistance outcomes. In addition to microbial community dynamics in the presence of antibiotics, I explore the interplay between tolerance and resistance that arises in anaerobic biofilms.
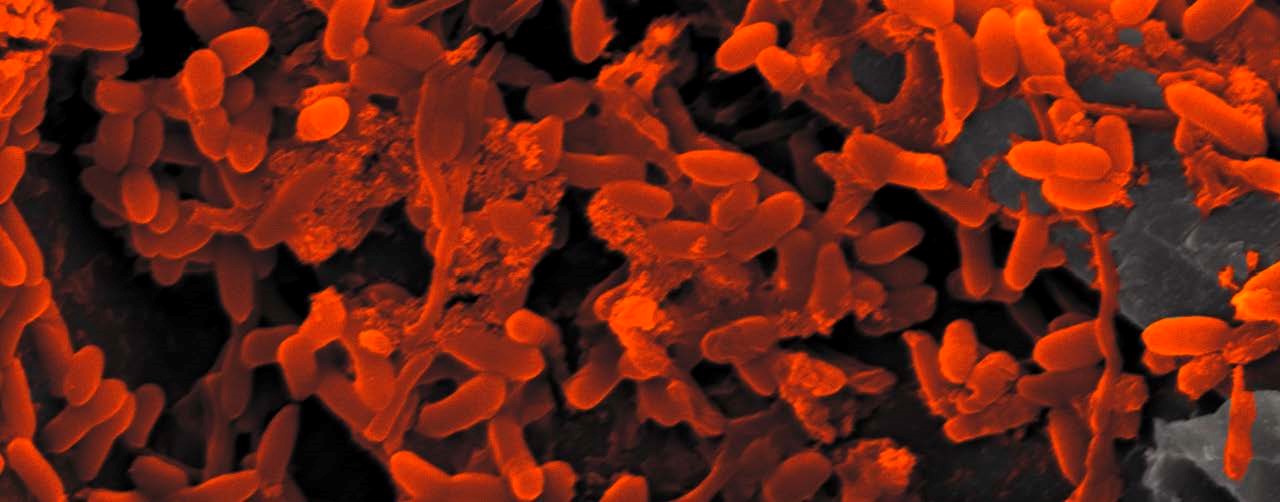 Life is the movement of electrons for the purpose of energy conservation. The scope of these electron transfer reactions in bacteria and archaea is beautifully eloquent and diverse. I use bioelectrochemical systems (electrodes) to monitor electron transfer reactions in biofilms. These systems allow for the real-time study of metabolic activity, disturbance challenges, diffusion limitations, and turnover of the microbial community.
Life is the movement of electrons for the purpose of energy conservation. The scope of these electron transfer reactions in bacteria and archaea is beautifully eloquent and diverse. I use bioelectrochemical systems (electrodes) to monitor electron transfer reactions in biofilms. These systems allow for the real-time study of metabolic activity, disturbance challenges, diffusion limitations, and turnover of the microbial community.
Scribner MR, Santos-Lopez A, Marshall CW, Deitrick C, Cooper VS. Parallel Evolution of Tobramycin Resistance Across Species and Environments. 2020. mBio. 11, e00932-20.
Santos-Lopez* A, Marshall* CW, Scribner MR, Snyder DJ, Cooper VS. Evolutionary Pathways to Antibiotic Resistance are Dependent Upon Environmental Structure and Bacterial Lifestyle. bioRxiv. 2019. https://doi.org/10.1101/581611.
Payne RB, Ghosh U, May HD, Marshall CW, Sowers KR. A Pilot-Scale Field Study: In Situ Treatment of PCB-Impacted Sediments with Bioamended Activated Carbon. Environmental Science and Technology. 2019. DOI: 10.1021/acs.est.8b05019
Gloag* ES, Marshall* CW, Snyder DJ, Lewin GR, Harris JS, Chaney SB, Whiteley M, Cooper VS, Wozniak DJ. The Pseudomonas aeruginosa Wsp pathway undergoes positive evolutionary selection during chronic infection. mBio. 2019. https://mbio.asm.org/content/10/4/e01698-19.
Dunlap DG, Marshall CW, Fitch A, Rapport SF, Cooper VS, McVerry B, Morris A, GD Kitsios. Bacterial DNA next generation sequencing can detect culprit pathogens missed by cultures and impact antibiotic management decisions in severe bacterial pneumonia. American Journal of Case Reports. 2018. 19, 1405-1409.
Turner C, Marshall CW, Cooper V. Parallel genetic adaptation across environments differing in mode of growth or resource availability. 2018. Evolution Letters. doi.org/10.1002/evl3.75.
Marshall CW, DE Ross, KM Handley, PB Weisenhorn, CS Henry, JA Gilbert, HD May, RS Norman. Metabolic Reconstruction and Modeling Microbial Electrosynthesis. 2017. Scientific Reports. 10.1038/s41598-017-08877-z (SREP-17-14408).
Marshall CW, EV LaBelle, HD May. Production of Fuels and Chemicals from Waste by Microbiomes. Current Opinions in Biotechnology. 2013. 24(3): 391-397.
Sophie Bridwell (Ph.D. student)
Marissa Allen (Ph.D. student)
Dr. Marshall is currently accepting new Ph.D. students into his lab